BinAC2/Software/Jupyterlab
|
The main documentation is available on the cluster via |
| Description | Content |
|---|---|
| module load | devel/jupyterlab |
| License | JupyterLab License |
| Links | Homepage |
| Graphical Interface | Yes |
Description
JupyterLab is a web-based interactive development environment for notebooks, code, and data.
Currently, BinAC 2 provides the following JupyterLab Docker images via Apptainer:
minimal-notebookr-notebookjulia-notebookscipy-notebook
Usage
This guide is valid for minimal-notebook. You can also follow it for the other notebook flavors — just replace minimal-notebook with one of the other notebooks listed above.
You will start a job on the cluster as usual, create an SSH tunnel, and connect to the running JupyterLab instance via your browser.
Start JupyterLab
The module devel/jupyterlab provides a job script for starting a JupyterLab instance on BinAC 2. Load the module and copy the template job script into your workspace:
$ module load devel/jupyterlabUpon loading the module, you can list the available template job scripts and copy the one you need into your workspace:
$ ls $JUPYTERLAB_EXA_DIR
binac2-julia-notebook.slurm binac2-minimal-notebook.slurm binac2-r-notebook.slurm binac2-scipy-notebook.slurm
$ cp $JUPYTERLAB_EXA_DIR/binac2-minimal-notebook.slurm <your workspace>The job script is very simple; you only need to adjust the hardware resources according to your needs:
#!/bin/bash
# Adjust these values as needed
#SBATCH --cpus-per-task=1
#SBATCH --mem=2gb
#SBATCH --time=6:00:00
# Don't change these settings
#SBATCH --nodes=1
#SBATCH --partition=interactive
#SBATCH --job-name=minimal-notebook-7.4.1
#PBS -j oe
# Load Jupyterlab module
module load devel/jupyterlab/7.4.1
# Start jupyterlab
$ ${JUPYTERLAB_BIN_DIR}/minimal-notebook.sh
Now submit the job to SLURM. This command will store the job ID in the variable jobid
$ jobid=$(sbatch --parsable binac2-minimal-notebook.slurm)|
Depending on the current load on BinAC 2 and the resources you requested in your job script, it may take some time for the job to start. |
Create SSH tunnel
The compute node on which JupyterLab is running is not directly reachable from your local workstation. Therefore, you need to create an SSH tunnel from your workstation to the compute node via the BinAC 2 login node.
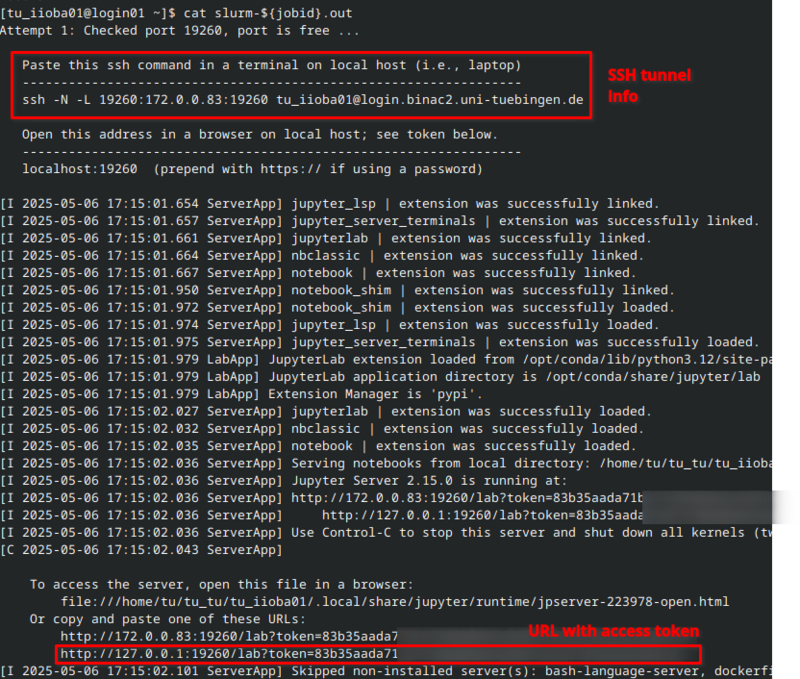
The job's standard output file (slurm-${jobid}.out) contains all the information you need. Please note that details such as the IP address, port number, and access URL will differ in your case.
$ cat slurm-${jobid}.out
Linux Users
Copy the ssh -N -L ... command and execute it in a shell on your workstation. After successful authentication, the SSH tunnel will be ready to use. The ssh command does not return any output. If there are no error messages, everything should be fine:
Windows Users
If you are using Windows, you will need to create the SSH tunnel using an SSH client of your choice (e.g. MobaXTerm, PuTTY, etc.). Her, we will show how to create an SSH tunnel with MobaXTerm.
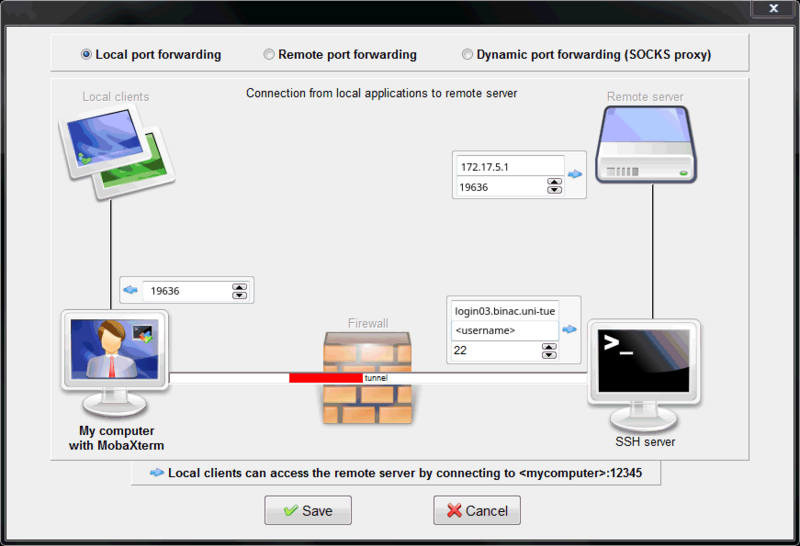
Select Tunneling in the top ribbon. Then click New SSH tunnel.
Configure the SSH tunnel with the correct values taken from the SSH tunnel information provided above.
For the example in this tutorial it looks as follows:
Access JupyterLab
JupyterLab is now running on a BinAC 2 compute node, and you have created an SSH tunnel from your workstation to that compute node. Open a browser and paste the URL with the access token into the address bar:
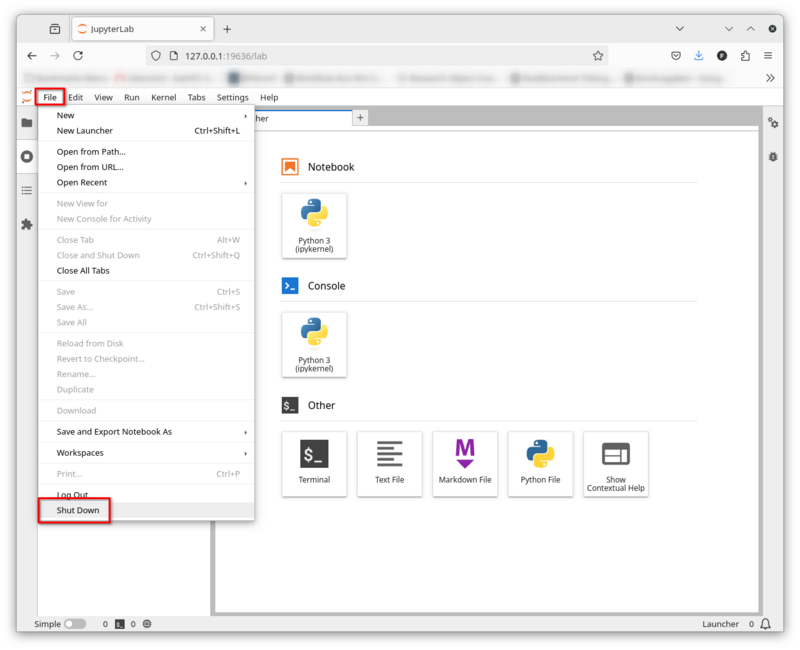
Your browser should now display the JupyterLab interface:
Shut Down JupyterLab
You can shut down JupyterLab via File -> Shut Down.
Please note that this will also terminate your compute job on BinAC 2!
Tips & Tricks
Managing Notebooks
JupyterLab's root directory will be your home directory. Since your home directory is backed up daily, you may want to store your notebooks there. It may also be a good idea to place your notebooks under proper version control using Git.
Access work and project
The notebooks will always be able to access your data stored in work and project. However, the file browser shows the content in your home directory and you won’t be able to access work and project initially.
You can create symbolic links in your home directory to work and project, making these two partitions available in JupyterLab's file browser.
$ cd ~
$ ln -s /pfs/10/work/ $HOME/work
$ ln -s /pfs/10/project $HOME/project
Through that link in your home directory you can navigate to your research data in Jupyterlab's file browser. Here is an example how I linked to my directory:
Managing Kernels
Depending on the notebook you started, the available kernels will differ.
The Python notebook, for example, only has one Python kernel, whereas the R notebook has only one R kernel, and so on.
Given the nearly endless combinations of programming languages and packages they provide, we suggest creating your own kernels if needed.
The kernels are stored in your hone directory on BinAC 2: $HOME/.local/share/jupyter/kernels/
You cannot install new kernels directly from within JupyterLab; this must be done from the command line in your usual BinAC 2 shell session.
Add a new Kernel
We will show you how to create new kernels for Python and R.
Python
module load devel/miniforge
conda create --name kernel_env python=3.13 pandas numpy matplotlib ipykernel # 1
conda activate kernel_env # 2
python -m ipykernel install --user --name pandas --display-name="Python 3.13 (pandas)" # 3
# Installed kernelspec pandas in $HOME/.local/share/jupyter/kernels/pandas
The first command creates a new Conda environment called kernel_env and installs a specific Python packages , along with a few additional Python packages.
It's important that you also install ipykernel, as we will need it later to create the JupyterLab kernel.
The second command activates the kernel_env Conda environment.
The third command creates the new JupyterLab kernel.
R
The instructions for creating new R kernels are slightly different.
module load devel/miniforge
conda create --name r_kernel_env r-base=4.4.3 jupyter r-irkernel
conda activate r_kernel_env
R
# In the R-Session
install.packages(...)
IRkernel::installspec(name = 'ir44', displayname = 'R 4.4.3')
The first command creates a new Conda environment called r_kernel_env and installs a specific R version.
It's important to also install r-irkernel, as we will need it later to create the JupyterLab kernel.
The second command activates the r_kernel_env Conda environment and open an R session.
In this session, you can install any R packages you need for your kernel.
Finally, create the new kernel with the installspec command.
Remove a Kernel
To remove a kernel from JupyterLab, simply delete the corresponding directory at $HOME/.local/share/jupyter/kernels/:
# Remove the JupyterLab kernel installed in the previous example
rm -rf $HOME/.local/share/jupyter/kernels/pandas
Also, remove the corresponding Conda environment if you do not need it any more:
conda env remove --name kernel_env